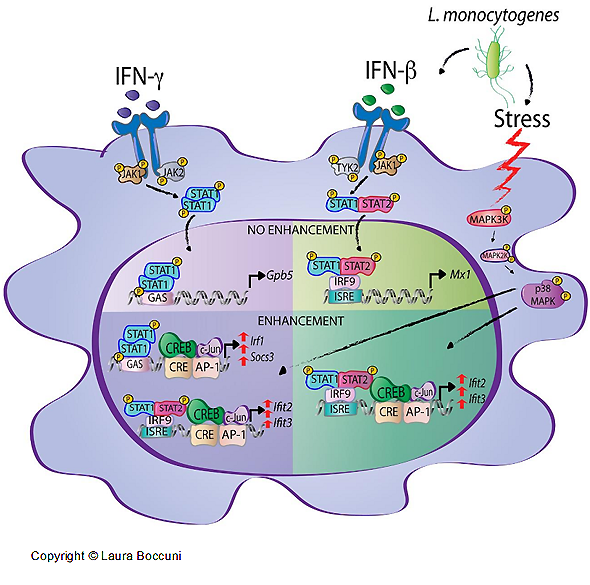
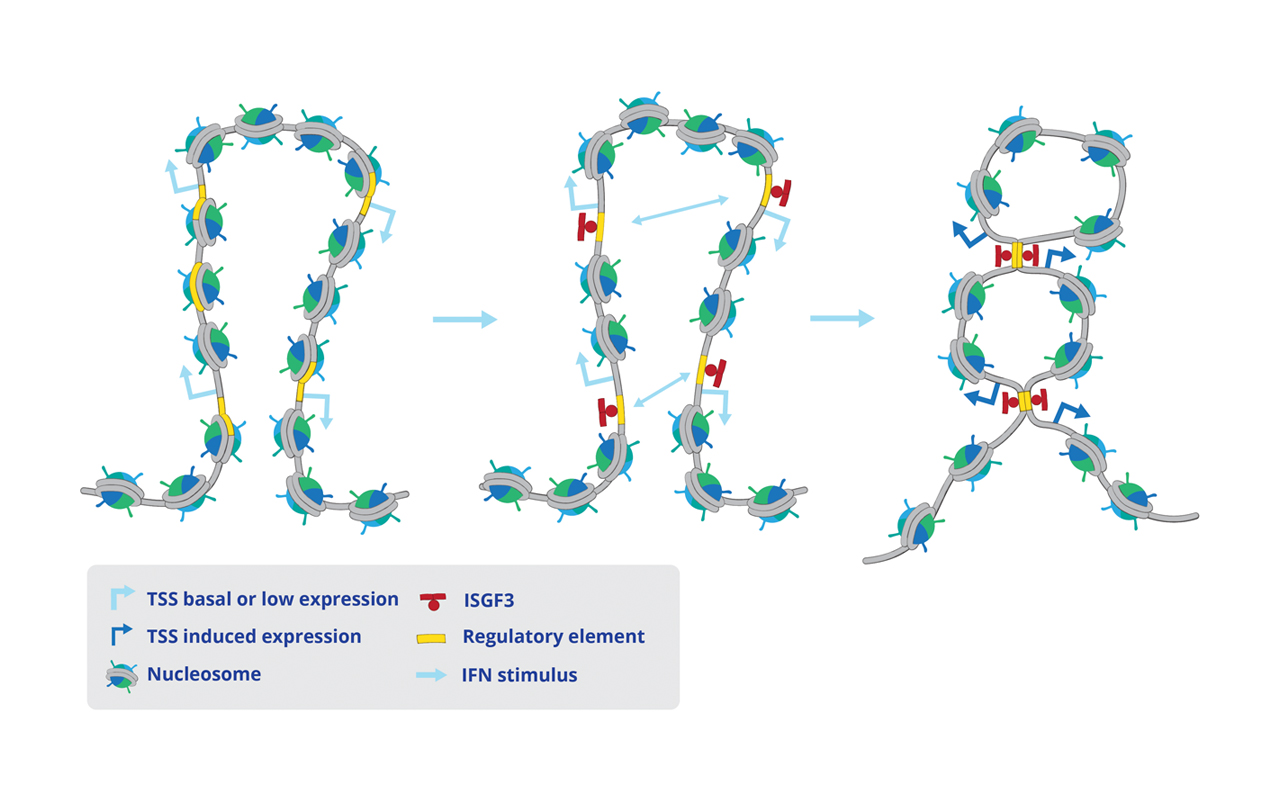
Interferons (IFNs) activate cell-intrinsic defence mechanisms to fight infections. The innate immune response to infections overlaps and cooperates with cellular stress signalling governed by the mitogen-activated protein kinase (MAPK) p38. Laura Boccuni in the group of Thomas Decker unravelled the molecular mechanism how macrophages merge the IFN JAK-STAT and the stress p38 MAPK pathways to enhance the beneficial but also the detrimental effects of the innate and inflammatory responses. With the exception of one, all SFB groups contributed to this SFB-financed publication.
Published in Science Signaling
Laura Boccuni, Elke Podgorschek, Moritz Schmiedeberg, Ekaterini Platanitis, Peter Traxler, Philipp Fischer, Alessia Schirripa, Philipp Novoszel, Angel R. Nebreda, J. Simon C. Arthur, Nikolaus Fortelny, Matthias Farlik, Veronika Sexl, Christoph Bock, Maria Sibilia, Pavel Kovarik, Mathias Müller, Thomas Decker
Stress signaling boosts interferon-induced gene transcription in macrophages
https://doi.org/10.1126/scisignal.abq5389
See also for RELATED FOCUS: Rewiring the logic board of IFN signaling, by John D. MacMicking John D. MacMicking
https://doi.org/10.1126/scisignal.adf0778